Finding faster, more accurate ways to test drugs using AI and machine learning
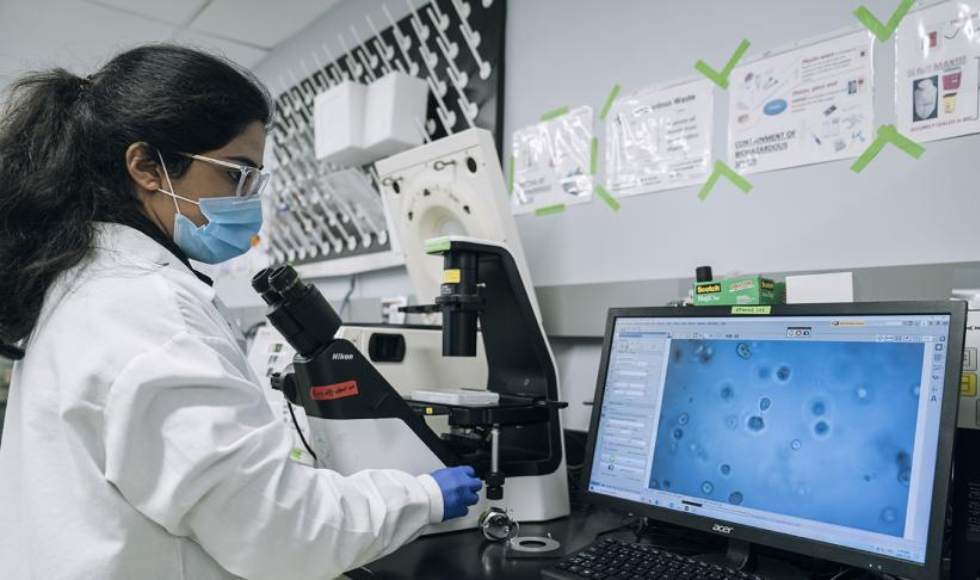

Photos by Jin Lee
BY Jessie Park, Faculty of Engineering
November 11, 2020
A team of student researchers at McMaster Engineering have applied machine learning and artificial intelligence (AI) to detect changes in the way human cells organize themselves in response to drug treatments – paving the way for faster and more accurate testing for drugs, such as medications to treat viruses and cancers.
By using AI to study the ability of cells to self-organize into their native structures – or what tissues they “should” form – the team’s approach shows results beyond the basic parameters of testing whether cells are alive or not after drug treatment.

“We were interested in using AI or deep learning to see if it could classify between two different tissue structures of lung cells when they’re grown in the lab,” says Lyan Abdul, a master’s student in Biomedical Engineering who began this research as a Life Sciences undergraduate student.
One organizational structure of lung alveolar cells – their native structure – is a sphere-like shape with a hollow centre, “like a donut,” says Abdul.
The second structure is more disorganized and more of a solid sphere, which indicates a non-native structure for lung alveolar cells.
“This is our first time applying machine learning to image analysis for drug testing,” says Boyang Zhang, an assistant professor of Chemical Engineering who supervised Abdul and the co-authors on a paper published in the journal Lab on a Chip.
“We created a data set with example images of these two structures, and trained the AI machine to analyze whether the tissue structure was hollow or solid,” he adds.
Eventually, the AI machine could detect the cell structures about 20 times faster than Abdul could do manually – and with more specific and non-biased results.
“With this method, we were able to uncover that at a certain dosage, a drug called cyclosporin affects the self-organization of lung cells in a significant way,” she says, emphasizing that without using AI, she would have had to use a fluorescent staining method to analyze the cell’s structures.
Cyclosporin is an immunosuppressant drug commonly used after organ transplants, she says.
“Fluorescent staining is damaging to the cells and time-consuming for the researcher because you have to stop the experiment each time to stain them,” Abdul adds. “By removing that process and using AI, the cells were analyzed in three minutes whereas it could take me an hour to do manually.”
This is just the beginning, according to Zhang, who leads the BZhangLab at McMaster dedicated to creating innovative biotech solutions to improve upon existing methods of healthcare.
“This is one specific application we’ve tested, and now with this machine learning capability in our lab we can expand it to many other applications on different types of tissues and organ systems,” he says.
“The next step is training the machine to recognize patterns that we cannot even recognize by eye – and that’s totally doable.”
Abdul, who previously received the NSERC Undergraduate Student Research Award for two summers in a row at McMaster, says this research was a huge team effort of undergraduate and graduate students.
The co-authors on the paper are graduate students Shravanthi Rajasekar and Dawn Lin, and undergraduate students Sibi Raja, Alexander Sotra, Yuhang Feng and Amy Liu.
Abdul’s advice for up-and-coming student researchers?
“Don’t limit yourself. In the beginning, I knew nothing at all about AI or machine learning – don’t let that stop you from getting involved in research.”